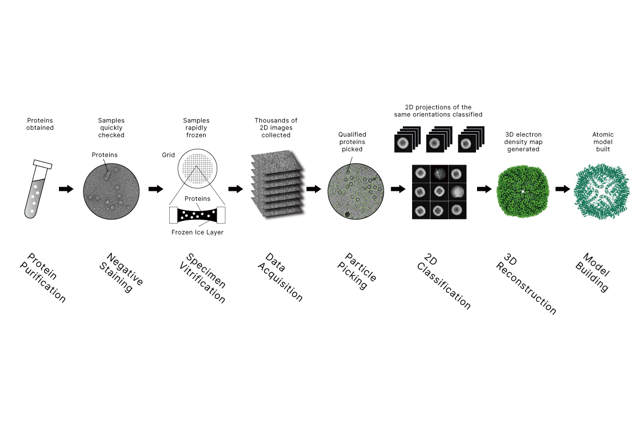
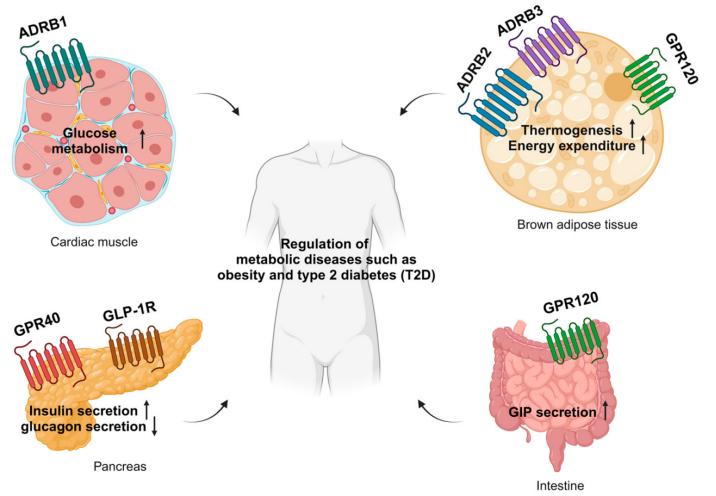
Related Post
Analyze Membrane Proteins with XL-MS: Step-by-Step Workflow Guide
2026-02-20Analyze Membrane Proteins with XL-MS is one of the most practical ways to turn “hard-to-see” membrane interactions into measurable evidence. Longlight Technology uses chemical cross-linking coupled with mass spectrometry (XL-MS) to help researchers map protein–protein interactions (PPIs), capture weak or short-lived contacts, and move from hypotheses to defensible structural models—without forcing special chemical labeling workflows.
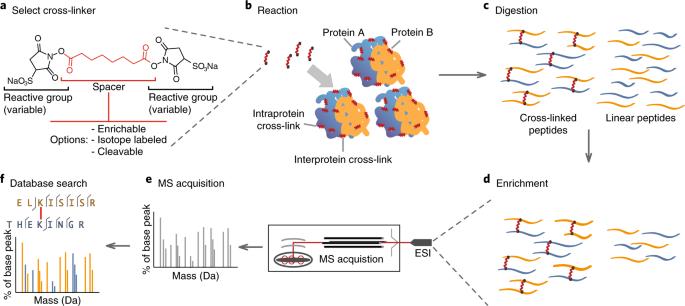
Cross-linking mass spectrometry: methods and applications in structural, molecular and systems
What Is XL-MS? What Are Membrane Proteins?
XL-MS and membrane proteins are two ideas that often show up together in structural biology and drug discovery. Here’s a clear, beginner-friendly explanation.
What Is XL-MS?
XL-MS stands for cross-linking mass spectrometry (often written as chemical cross-linking coupled with mass spectrometry).
It is a method used to study protein–protein interactions and protein shapes by doing two things:
Cross-linking (XL): “Freeze” nearby parts in place
A chemical cross-linker acts like a tiny molecular “bridge.” If two amino-acid sites are close enough in real space (within a short distance), the cross-linker can covalently connect them.
This helps preserve weak or short-lived interactions that might fall apart during normal sample handling.
Mass Spectrometry (MS): Identify the linked pieces
After cross-linking, proteins are digested into peptides. The mass spectrometer detects cross-linked peptide pairs, and software maps them back to proteins.
Those links provide distance clues (spatial restraints), telling you which regions were close in the original complex.
What XL-MS tells you (in practice):
• Which proteins likely interact (interaction network)
• Which regions touch (interface hints)
• Structural constraints that support models (often combined with cryo-EM or X-ray)
What Are Membrane Proteins?
Membrane proteins are proteins that are embedded in or attached to a cell membrane (or membranes inside cells, like the ER, mitochondria, or bacteria membranes).
They are critical because membranes control transport and signaling, and membrane proteins often act as “gatekeepers” or “antennas” for the cell.
Main Types
• Integral membrane proteins: physically embedded in the membrane
• Many span the membrane one or multiple times (often called transmembrane proteins)
• Peripheral membrane proteins: attached to the membrane surface (often via other proteins or lipids)
Why Membrane Proteins Matter
• They are involved in:
• Transport (channels, pumps, transporters)
• Cell signaling (receptors like GPCRs)
• Energy conversion (respiratory chain complexes)
• Cell recognition and adhesion
Why They Are “Hard”
Membrane proteins are challenging to study because:
• They sit in a lipid environment
• They can be unstable when removed from the membrane
• They often form dynamic complexes that change states
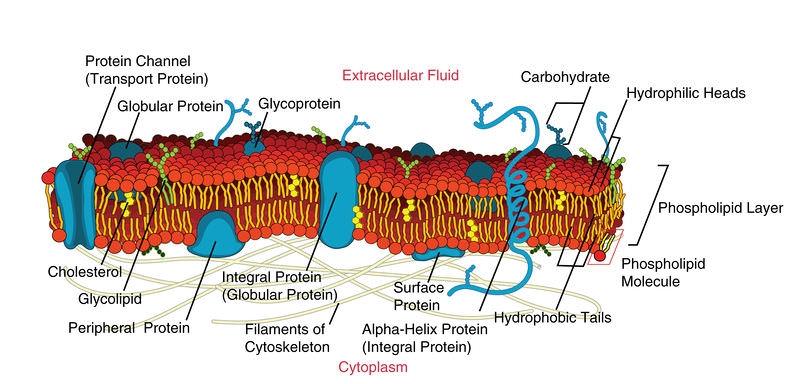
Membrane Proteins - Biology LibreTexts
Why Membrane Proteins Need a Different Strategy
Membrane proteins often sit in crowded, dynamic environments. Their key interfaces can be transient, and many complexes are unstable once removed from the membrane. That is why classic interaction methods may miss critical contacts or over-simplify what is happening.
Analyze Membrane Proteins with XL-MS works because it “freezes” nearby regions in place using cross-linking agents. These reagents can covalently connect two or more interacting proteins within a defined distance window. Once the interaction is locked in, mass spectrometry can read out cross-linked peptides and point you to likely contact sites. Operationally, interaction evidence remains traceable even when complexes are weak.
• Secures short-lived interactions often lost during purification
• Avoids chemical labeling, simplifying early decisions
• Enables intracellular cross-linking to preserve native proximity
Step 1: Specify the biological question and distance rationale
Decide what you want to learn before you begin. Are you confirming a suspected partner? Comparing mutant vs wild type binding? Mapping an interface to support a model? A clear question helps you choose conditions, controls, and data outputs.
XL-MS is distance-informed. Cross-linking agents connect residues that are close in space, not necessarily adjacent in sequence. For membrane proteins, this distance logic is valuable because it can reveal how helices pack, how cytosolic loops contact partners, or how oligomers assemble.
A beginner-friendly way to frame your plan is to define three outputs:
• Interaction Presence: does A contact B at all?
• Interaction Topology: which regions of A contact which regions of B?
• Site Evidence: which peptide pairs support the interaction network?
Analyze Membrane Proteins with XL-MS becomes much smoother when you pre-decide which of these you need for your next decision (structure modeling, target validation, or mechanism study).
Step 2: Choose a Cross-Linking Approach That Fits Membrane Context
Membrane proteins are sensitive to detergents, lipid mimetics, and buffer composition. The goal is to maintain a state that resembles the functional complex as closely as possible, then cross-link at the right moment.
At Longlight Technology, customers can either send already cross-linked samples or contact us to develop a cross-linking plan and then submit samples. This flexibility matters for membrane targets because one project may require in-solution cross-linking, while another benefits from more native-like conditions or even intracellular cross-linking.
✔ High throughput and fast analysis speed can shorten iteration cycles
✔ Intracellular cross-linking can reduce artifacts from over-handling complexes
✔ No special labeling requirement keeps early experiments accessible
Practical tip for beginners: plan at least one negative control (no cross-linker, or no partner protein) and one “known behavior” reference if available. Controls help you distinguish real proximity signals from background.
Step 3: From Cross-Linked Proteins to Detectable Peptides
After cross-linking, the workflow must translate proteins into a peptide mixture that still preserves the cross-link information. This is where many beginners feel lost, because cross-linked peptides are rarer than normal peptides and can be harder to detect.
Our standard service workflow covers the full chain:
• Enzyme digestion
• Peptide enrichment
• Mass spectrometry detection
• Data analysis
• Experimental report delivery
The key idea is simple: digestion turns the proteins into peptides, enrichment increases the relative visibility of cross-linked species, and mass spectrometry measures the peptide masses and fragments so the software can assign cross-link pairs.
Analyze Membrane Proteins with XL-MS is most efficient when you treat sample prep as an information-preservation step, not just a routine protocol. The best results come from stable digestion and careful enrichment so the data carries enough signal for confident interpretation.
Step 4: Mass Spectrometry Readout and Interaction Network Mapping
Mass spectrometry does more than “identify proteins.” In XL-MS, it identifies cross-linked peptide pairs, which you can interpret as spatial restraints. Once you have enough high-confidence links, you can begin to depict interaction networks and infer sites of action—especially useful for complex membrane assemblies.
This is also where XL-MS becomes a bridge to structural biology. Many teams use XL-MS outputs alongside cryo-EM or X-ray crystallography. The cross-link evidence can help:
• Validate whether a structural model is plausible
• Resolve ambiguous subunit orientations
• Support domain placement when density is limited
In other words, Analyze Membrane Proteins with XL-MS can move you from “we think these two regions interact” to “we have distance-supported evidence that constrains the model.”
Step 5: How to Read the Report and Turn Data Into Next Experiments
A report is only valuable if it guides decisions. For beginners, the most useful way to read XL-MS results is to look for patterns, not just lists.
Start with three questions:
✔ Are cross-links reproducible across replicates or conditions? Reproducibility builds confidence.
✔ Do the links cluster in specific regions? Clustering often points to real interfaces.
✔ Do mutant/ligand conditions shift the link pattern? Shifts can reveal mechanism.
Then translate evidence into next steps. If links support a specific interface, you can design point mutations for validation. If links suggest unexpected partners, you can plan orthogonal confirmation. If links constrain a model, you can proceed to structural refinement with higher confidence.
CTA: If you want to Analyze Membrane Proteins with XL-MS but are unsure how to choose conditions or controls, contact Longlight Technology to develop a cross-linking plan and receive a complete workflow report—from digestion to data analysis. Get a free quote to start your project with a clear, beginner-friendly experimental roadmap.
Why Longlight Technology for XL-MS and Beyond
Membrane protein projects rarely live in a single technique. They typically require a platform mindset: reliable sample handling, rigorous data, and tools that accelerate iteration.
Longlight Technology provides all-round support with cutting-edge genomics solutions, advanced laboratory instruments, and high-quality reagents and consumables designed to improve efficiency and accuracy in modern laboratories. In addition to XL-MS services, we support research teams with genomics and molecular biology tools, including NGS-related instruments such as focused ultrasonication systems, plus widely used consumables and kits (precast agarose gels, nucleic acid extraction kits, and library preparation kits) for academic, clinical, and industrial applications.
Analyze Membrane Proteins with XL-MS is not only a method—it is a workflow discipline. When the workflow is stable, your conclusions become clearer, your models become more defensible, and your next experiment becomes easier to design.