Related Post
Choosing the Right Expression Platform for XL-MS Membrane Proteins
2026-02-19Choosing the Right Expression Platform for XL-MS Membrane Proteins starts with one reality: membrane targets are the most valuable—and often the most difficult—proteins you will ever bring into a structural workflow. They sit inside lipid bilayers, shape transport and signaling, and dominate modern drug discovery. In fact, "membranome" networks account for roughly 30% of the mammalian proteome and about 60% of drug targets, which is why robust membrane-protein workflows matter so much.
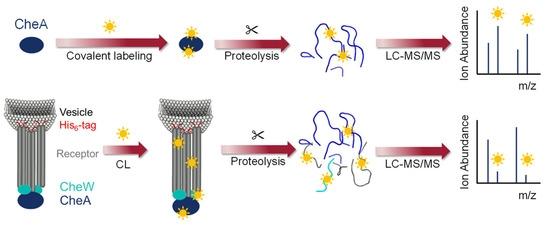
Advances in Mass Spectrometry on Membrane Proteins
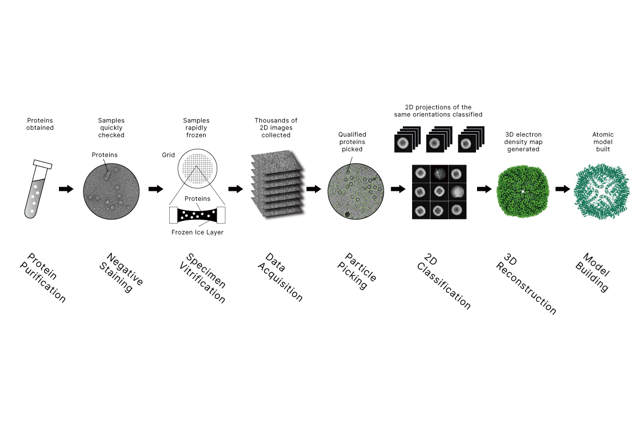
At Longlight Technology, we support XL-MS (chemical cross-linking coupled with mass spectrometry) projects designed to map protein–protein interactions (PPIs) and capture short-lived or weak contacts through covalent cross-links. XL-MS has become a core method in integrative structural biology and is frequently combined with cryo-EM and other techniques when you need both interaction evidence and 3D context.
What Are "XL-MS Membrane Proteins"?
What XL-MS Means
XL-MS = chemical cross-linking coupled with mass spectrometry.
Cross-linking (XL): A small chemical "linker" reacts with two amino acids that are close in space (typically within a limited distance). It forms a covalent bridge between them.
Mass spectrometry (MS): After digestion into peptides, MS detects which peptide pairs are cross-linked, letting you infer who is near whom and which regions are close.
What "Membrane Proteins" Are
Membrane proteins are proteins embedded in or attached to cell membranes (e.g., GPCRs, ion channels, transporters, receptors). They often:
• Have hydrophobic transmembrane segments
• Require lipids/detergents to stay stable outside the membrane
• Form complexes with other proteins
So What Are "XL-MS Membrane Proteins"?
It means using XL-MS to learn structure and interactions of membrane proteins, such as:
✅ Protein–protein interactions (PPIs): which partners bind to the membrane protein
✅ Interface mapping: which regions of the proteins contact each other
✅ Distance restraints: approximate spatial constraints that help model the protein/complex shape
✅ Capturing weak/transient interactions: cross-linking can "freeze" interactions that might fall apart during purification
✅ Supporting integrative structure work: often combined with cryo-EM or modeling to refine complexes
A Simple Example
If two parts of a membrane receptor (or the receptor + a partner protein) are close enough, the cross-linker connects them. MS then identifies the linked peptide pair. That tells you:
"These two residues were near each other in the sample’s 3D state."
XL-MS Common Application Field
1) Mapping GPCR Signaling Complexes (Drug-Target Receptors)
GPCRs (G-protein–coupled receptors) are classic membrane drug targets, but they are dynamic and hard to "freeze" in one shape.
How XL-MS is used
• Cross-links capture which parts of the GPCR sit near G proteins or other partners in an activated state.
• Those distance constraints help build integrative structural models, often alongside cryo-EM.
Famous example
• XL-MS + integrative modeling was used to map the conformational ensemble of an activated GLP-1 receptor–Gs complex (important in metabolic disease research).
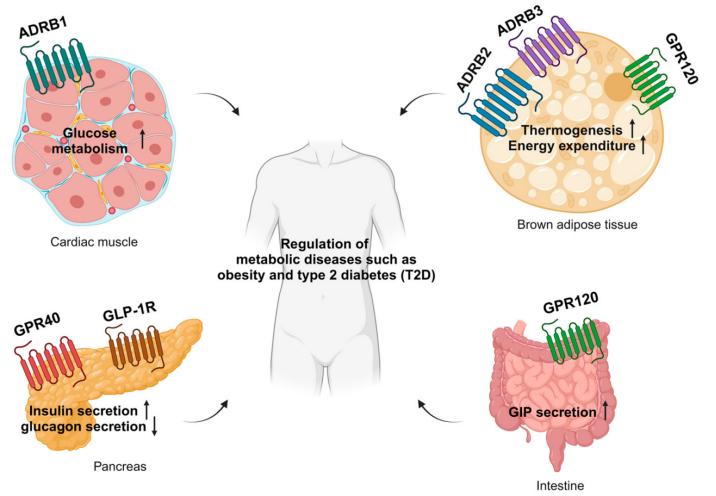
G-Protein-Coupled Receptor (GPCR) Signaling and Pharmacology in Metabolism
2) Revealing Membrane Protein Interaction Networks In Organelles (Mitochondria)
Mitochondria contain many membrane complexes (respiratory chain complexes, transporters). XL-MS has been used to map how these proteins organize and interact in their native organelle environment.
How XL-MS is used
• Cross-linking can be done on intact mitochondria to preserve native contacts.
• MS identifies many residue-to-residue contacts → builds interaction networks.
Famous example
• "Interactome of intact mitochondria" studies used XL-MS to provide large-scale interaction maps and evidence related to respiratory supercomplex organization.
3) Capturing Weak/Transient Interactions That Purification Loses
A big reason XL-MS became popular is its ability to covalently lock interactions that are weak, transient, or short-lived—common for membrane assemblies.
Why it matters
• Many membrane complexes fall apart in detergent or during enrichment.
• XL-MS can "freeze" contacts early, so you don’t lose key partners.
This capability is highlighted in major XL-MS reviews and platform descriptions.
4) Integrative Structural Biology With Cryo-EM / Cryo-ET
For membrane proteins, cryo-EM may give you an overall shape, but flexible regions or subunit placement can remain uncertain. XL-MS provides distance restraints that help:
• Position subunits,
• Validate interfaces,
• Constrain flexible regions.
This "XL-MS + cryo methods" combination is a mainstream integrative workflow.
Why Expression Choice Is the First XL-MS Decision
Membrane proteins fail in XL-MS for predictable reasons: misfolding, incorrect oligomerization, missing post-translational modifications (PTMs), or harsh solubilization that destroys native contacts before cross-linking begins. Your expression platform quietly controls all of these variables.
For beginners, a helpful mental model is this: XL-MS does not "fix" protein quality—it reports what you actually made. Reviews and method papers consistently emphasize that XL-MS becomes most informative when the starting material preserves native assemblies, whether in vitro or in situ.
So the right question is not "Which platform gives the highest yield?" but "Which platform gives the most biologically faithful sample for my cross-linking goal?"
What "Good Enough" Looks Like for XL-MS Membrane Proteins
Before choosing a host, define your success criteria in plain language. A membrane protein that is "expressed" is not automatically "XL-MS ready."
Here are practical benchmarks we recommend:
✓ Native-like state: correct topology, stable complex assembly, and consistent behavior during solubilization or reconstitution
✓ Cross-linkable environment: compatible buffer, mild detergents (or lipid mimetics), and minimal interfering additives
✓ Reproducible batch quality: similar yield and purity across repeats, so cross-links reflect biology—not batch drift
✓ Right level of complexity: purified complex when you need precise topology; intracellular/in situ when you need native context and transient partners
XL-MS is especially valued because cross-linkers can "freeze" weak or short-lived interactions that are otherwise lost during purification. That advantage only shows up when your expression system helps maintain those interactions long enough to capture them.
A Beginner-Friendly Map of Expression Platforms
Different platforms solve different failure modes. Use this map to narrow your first choice.
E. coli And Yeast: Best For Speed, Screening, And Simple Targets
Bacteria and yeast can be excellent when your membrane protein is small, relatively stable, and not heavily PTM-dependent.
✓ Fast build–test cycles for constructs, truncations, and tags
✓ Cost-effective scale-up for purified samples
✓ Good for initial feasibility screens before moving to higher-fidelity hosts
Where they struggle is equally consistent: complex multi-pass eukaryotic receptors, fragile assemblies, and proteins that require mammalian PTMs for correct folding or partner binding.
Insect and Mammalian Cells: Best for Eukaryotic Folding and Native Assemblies
If your target depends on chaperones, glycosylation, or native complex formation, insect and mammalian systems often reduce the "looks fine on SDS-PAGE, fails in biology" problem.
✓ Higher chance of correct folding for GPCRs, channels, transporters, and receptors
✓ Better support for native-like complexes and functional conformations
✓ More suitable for pairing XL-MS with cryo-EM when you need integrative structure building
The tradeoff is time and cost. But for membrane-protein XL-MS, higher fidelity frequently saves weeks of downstream troubleshooting.
Match the Platform to Your XL-MS Goal
Many teams choose an expression system without stating the XL-MS "mode." We recommend deciding between two common modes up front.
Mode A: Purified/Enriched Complex XL-MS (High Interpretability)
You want clean identification of cross-linked peptides and confident interaction maps.
✓ Choose a platform that yields stable, enrichable complexes (often insect/mammalian for eukaryotes)
✓ Aim for mild solubilization, minimal aggregation, and consistent oligomer state
✓ Consider reconstitution (nanodiscs or lipid mimetics) when detergent destabilizes contacts
Mode B: Intracellular Or Near-Native XL-MS (High Biological Relevance)
You want native partners, transient contacts, and real cellular context. XL-MS literature highlights the growing frontier of in situ workflows, even though they remain technically demanding.
✓ Choose a platform that supports physiological localization and native binding partners
✓ Design cross-linking conditions to avoid over-crosslinking and non-specific networks
✓ Expect more complex data—but also more biologically meaningful interaction discovery
At Longlight Technology, we can work with either mode because our workflow supports enzyme digestion, peptide enrichment, mass spectrometry detection, and data analysis from a defined plan through to a final report.
Longlight XL-MS Service Advantages in Practical Terms
Our service advantages matter only if they translate into outcomes you can feel in a real project:
✓ High Throughput And Fast Analysis Speed → faster iteration when you are optimizing constructs, detergents, or host systems
✓ Intracellular Cross-Linking Capability → better chance to capture weak or transient partners before purification disrupts them
✓ No Special Protein Labeling Required → simpler sample preparation and fewer variables for first-time users
✓ Covalent Capture Of Short-Lived/Weak Interactions → stronger evidence for interaction networks and contact regions, especially for dynamic membrane complexes
We also build broader laboratory support around these workflows. Longlight provides cutting-edge genomics solutions and laboratory equipment designed to improve efficiency and accuracy in modern research—ranging from NGS-related instruments (including focused ultrasonication) to high-quality reagents, consumables, and library prep kits that fit upstream and adjacent workflows.
A Clear Service Process and the Next Step
A smooth XL-MS project is usually the result of one good planning call and a disciplined sample pathway.
Our Standard Process
• You can send cross-linked samples, or contact us to co-develop a cross-linking plan
• We complete enzyme digestion, peptide enrichment, MS detection, and data analysis
• We deliver a structured experimental report, including interaction/network interpretation and identified sites of action
If you are currently choosing the right expression platform and feel uncertain, start with the simplest decision support: tell us your target class (channel, transporter, receptor), the organism, and whether you want purified-complex XL-MS or intracellular XL-MS. We will help you align expression, solubilization, and cross-linking into one coherent workflow—so your first dataset is informative, not just "technically successful."
CTA: If you want a practical recommendation for your membrane protein (host system + sample format + XL-MS mode), contact Longlight Technology for a free project assessment and quote tailored to your target and timeline.