Related Post
What Cross-Linking MS for Protein Complexes Teach About Conformational Change
2026-02-22Cross-Linking MS for Protein Complexes is one of the most practical ways to "freeze" real protein contacts in place and then read them out by mass spectrometry—so you can learn how a complex changes shape, not just what it looks like in one snapshot. At Longlight Technology, we see many teams start with a simple question: Is my protein complex static, or is it switching between conformations that matter for function? This article explains, in a beginner-friendly way, what chemical cross-linking coupled with mass spectrometry (often called CL-MS or XL-MS) can teach you about conformational change, and how to turn results into decisions you can act on.
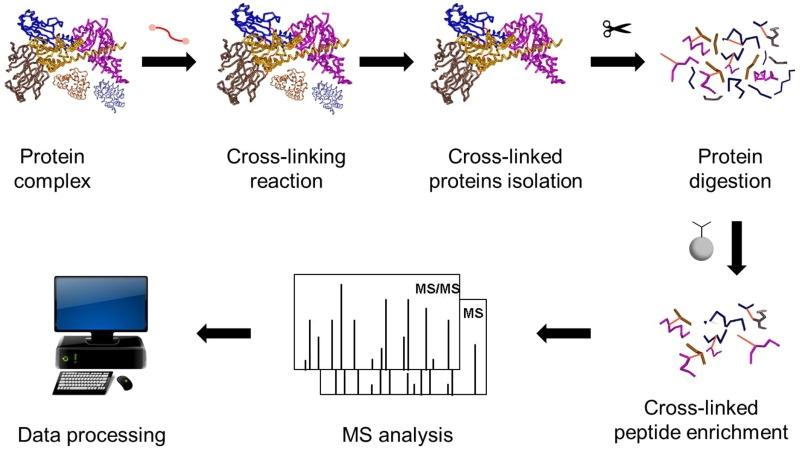
Advances in protein complex analysis by chemical cross-linking coupled with mass spectrometry
1) What is Cross-Linking MS?
Cross-Linking MS (often written XL-MS or CL-MS) is a method that helps you find which parts of proteins are close to each other—either within one protein or between proteins in a complex—by chemically "linking" them and then identifying those links by mass spectrometry.
Here’s the idea in plain English:
• Add a cross-linker (a small chemical "bridge")
It reacts with specific amino acids and can covalently connect two residues that are within a limited distance of each other.
• Digest the proteins into peptides
Enzymes (often trypsin) cut the proteins into smaller pieces.
• Run mass spectrometry
MS detects peptides, including cross-linked peptide pairs.
• Analyze the cross-links
Each identified cross-link becomes a distance constraint:
"Residue A and residue B were close enough to be linked under these conditions."
What it’s used for
• Protein–protein interaction (PPI) mapping: who touches whom in a complex
• Interface identification: which regions form the contact surface
• Conformational change: compare conditions (apo vs ligand-bound, mutant vs WT) to see contacts appear/disappear
• Supporting structural modeling: combine with cryo-EM/X-ray to validate or refine models
Why it’s valuable
• Can capture weak or short-lived interactions (the covalent link "freezes" them)
• Often no special labeling is required
• Can be relatively high-throughput for comparing multiple conditions
2) Why Conformational Change Is So Hard To Capture
Many protein complexes do not sit still. They breathe, pivot, open, close, and re-arrange subunits in response to ligands, salt, pH, phosphorylation, or binding partners. Traditional structural methods can be excellent, but they often favor stable states. If a complex is flexible, weakly assembled, or short-lived, you may only see part of the story.
Cross-linking helps because it can covalently connect two residues that come within a certain distance. In plain language, it marks "these two positions were close enough to touch" at the time of reaction. Side-by-side cross-link comparisons—ligand-free vs bound, low vs high salt, wild-type vs mutant—tell you if the complex compacts, expands, or reconfigures.
✅ Practical insight for beginners: conformational shifts are ensemble shifts. XL-MS sees beyond the most stable conformation to the spectrum of states in solution.
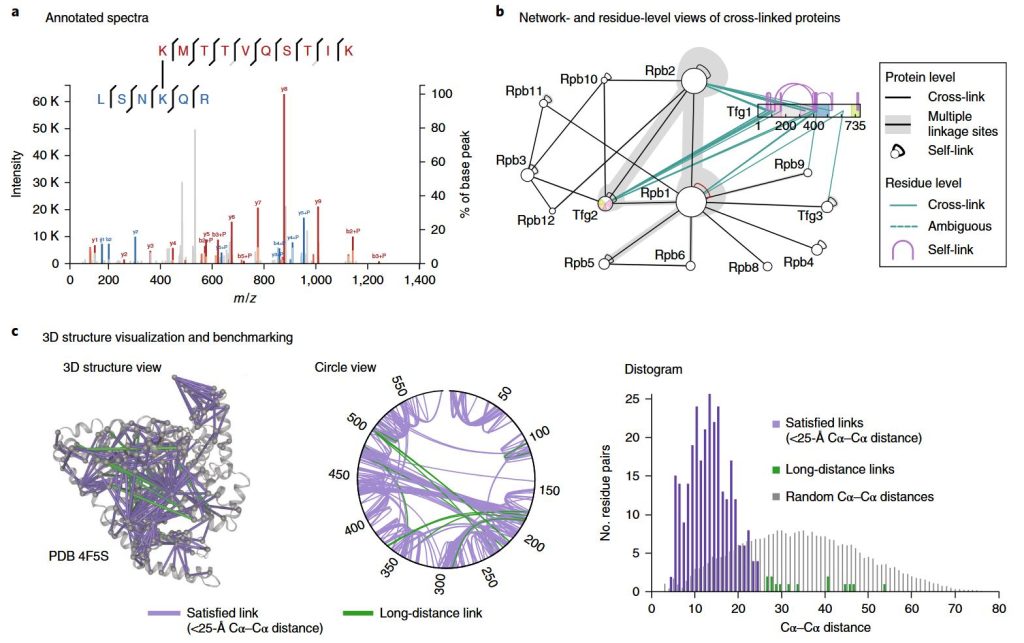
Crosslinking Mass Spectrometry Service | MtoZ Biolabs
What Cross-Linking MS Measures in Practice
Chemical cross-linking with MS quantifies residue proximity and interaction patterns, a standard approach for studying PPIs. Cross-linking agents react with functional groups on proteins and can connect two or more interacting proteins (or two positions within one protein). After cross-linking, mass spectrometry analyzes cross-linked peptides, allowing you to depict interaction networks and identify sites of action.
What does that mean for conformational change?
• If a set of cross-links appears only after ligand binding, it suggests new contacts formed in the bound state.
• If certain cross-links disappear, it suggests those sites are no longer close—perhaps the complex opens or a domain moves.
• If cross-links shift between subunits, it may indicate subunit rearrangement or a different assembly pathway.
✅ What you gain from this method (and why it matters):
• No special chemical labeling needed → you can keep your protein close to its native form and reduce experimental overhead.
• Captures short-lived/weak interactions → covalent bonds can preserve contacts that otherwise fall apart during purification or analysis.
• High throughput and fast analysis speed → useful when you need to compare many conditions or constructs efficiently.
• Intracellular cross-linking is possible → for some projects, this helps you study complexes closer to their native cellular context rather than only in vitro.
4) Reading "Motion" From Cross-Link Patterns
Beginners sometimes expect one cross-link equals one answer. In practice, the value comes from patterns.
A helpful way to think is: cross-links are distance constraints. When a complex changes conformation, the distance between two residues changes. XL-MS cannot always tell you the exact angle of rotation, but it can tell you whether regions likely moved closer or farther apart, and whether the interaction map changed.
Here are common conformational stories XL-MS can reveal:
✅ Compaction vs Opening
If you see more intra-protein cross-links spanning distant regions under one condition, the protein may be adopting a more compact state. If those links drop while others rise, it may be opening.
✅ Interface Switching
If cross-links between subunit A and B weaken, while links between A and C strengthen, it suggests a reweighted assembly or interface switch.
✅ Stabilization By Ligand Or Mutation
A ligand that "locks" a conformation often increases the reproducibility of a specific cross-link set and reduces mixed patterns.
From a practical standpoint, this can guide next steps: which mutant to make, which domain to truncate, which buffer condition stabilizes the complex, or which interface to validate by another method.
5) Stronger Results When XL-MS Is Combined With Cryo-EM Or X-Ray
XL-MS is often used alongside cryo-electron microscopy (cryo-EM) and X-ray crystal diffraction for biological structure research. The combination is especially useful when conformational change is the core question.
• Cryo-EM can provide a structural model for dominant states.
• XL-MS can validate whether a model is consistent with solution behavior and can flag alternative states that cryo-EM may under-sample.
• X-ray can deliver high-resolution domains, while XL-MS helps place domains within a flexible assembly.
✅ A practical workflow: use XL-MS first to learn whether your complex is heterogeneous. If it is, you can design conditions that enrich one state before investing heavily in high-resolution structure work.
6) The Service Workflow At Longlight Technology
Many labs want the insights of Cross-Linking MS for Protein Complexes without building a full pipeline in-house. Longlight Technology supports both experienced teams and first-time users with a clear service process.
You can either send cross-linked samples, or contact us to develop a cross-linking plan and then submit samples. We complete the full workflow, including enzyme digestion, peptide enrichment, mass spectrometry detection, data analysis, and delivery of an experimental report. This end-to-end approach matters because conformational interpretation depends on consistent handling across steps.
✅ What this means for you as a customer:
• Fewer handoff errors between steps, and fewer "unknowns" when you compare conditions
• A report that is organized around actionable interpretation, not only raw identifications
• Faster iteration when you need to test multiple constructs or treatment conditions
If your broader project includes genomics or upstream assay development, Longlight also provides cutting-edge genomics solutions, advanced laboratory instruments, and high-quality reagents and consumables designed to improve efficiency and accuracy in modern labs—supporting research workflows from molecular biology to precision analysis.
7) A Practical CTA: Turn Conformational Questions Into Testable Evidence
Conformational change is not a side detail. It often decides whether a target is druggable, whether a complex assembles correctly, and whether a mutation is truly disruptive. Cross-Linking MS for Protein Complexes gives you evidence you can compare across conditions, which helps you stop guessing and start designing.
✅ If you are planning a conformational-change study, consider starting with one "comparison set":
• Apo vs ligand-bound (or inhibitor-bound)
• Wild-type vs one interface mutant
• One stabilizing buffer vs one stressing buffer (salt/pH range)
CTA: If you want a clear, beginner-friendly XL-MS plan tailored to your protein complex, contact Longlight Technology to discuss your goal (interaction mapping, interface validation, or conformational comparison). We can help you choose a practical cross-linking strategy and deliver an interpretable report that supports your next experiment—or your next structural model.