Related Post
How To Validate Results: ChIP-Seq Data Analysis Pipeline Explained
2026-02-27ChIP-Seq Data Analysis Pipeline validation is the step that turns peaks into proof, not just pictures. At Longlight Technology, we support ChIP-seq projects from sample to report, and we see one clear rule: if you validate in the right order, your conclusions become stable, explainable, and easier to publish.
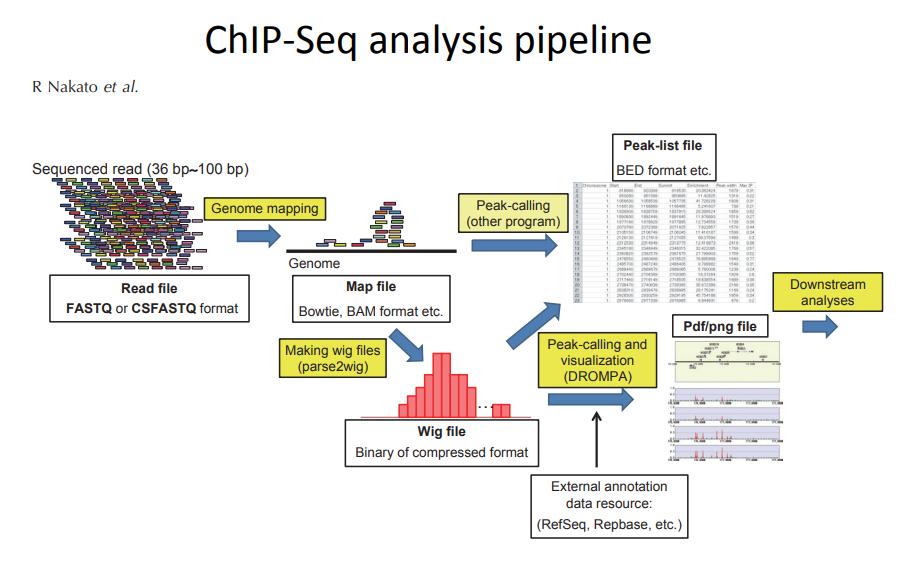
A Beginner's Guide to ChIP-Seq Data Analysis | by sezer islambey | Medium
1) Understand What ChIP-Seq Proves, and What It Does Not
ChIP-seq starts with a simple scientific goal: measure how proteins interact with DNA in real biological samples. In practice, it helps you map where transcription factors, RNA polymerase II, or histone modifications are enriched across the genome. A well-built ChIP-Seq Data Analysis Pipeline can support questions such as protein presence at different genomic sites, transcription factor binding behavior, and how histone marks relate to gene expression.
Validation matters because ChIP-seq is a chain of linked steps. If any one link is weak—antibody specificity, chromatin quality, library complexity, or background control—your final peaks may look convincing but still be unreliable. A beginner-friendly way to think about validation is this: you are not trying to “get peaks.” You are trying to show that the peaks would appear again under the same conditions, and that they match the biology you expect.
✓ ChIP-seq measures enrichment, not absolute binding counts
✓ Good results require controls and repeatability, not just sequencing depth
✓ Validation should be planned before the experiment starts, not after peaks are called
2) Build Validation Into the Workflow From the First Sample
Many teams only focus on validation at the bioinformatics stage. That is often too late. The most efficient validation approach is to make it part of the service process from the beginning, because early control prevents late-stage rework.
Outsource the entire pipeline: you provide fixed cell or frozen tissue samples; Longlight Technology delivers prep, chromatin processing, library construction, sequencing, and analysis. The benefit is not just convenience. It reduces handoff risk and keeps quality checks consistent across steps, which makes the final ChIP-Seq Data Analysis Pipeline output easier to trust.
When sample input is limited, validation becomes even more important. Our optimized process is suitable for small sample sizes, which helps researchers move forward when material is precious. The practical value for customers is simple: less sample waste, fewer trial-and-error repeats, and clearer decisions earlier in the project.
✓ One process owner reduces variation between steps
✓ Standard checkpoints prevent “invisible failures”
✓ Small-sample readiness protects rare tissues and low-yield projects
3) Validate Data Quality Before You Trust Any Peak
A strong ChIP-Seq Data Analysis Pipeline begins with quality control that answers one question: is your dataset capable of producing a real signal above background?
Start by reviewing off-machine data quality and basic library behavior. If the input DNA or ChIP library shows abnormal read quality, strong adapter presence, or extreme duplication, peak calling may amplify noise. Good validation does not chase perfection. It looks for consistency across samples and clear separation between ChIP and control.
Next, confirm alignment behavior and filtering rules. Mapping rate and the fraction of usable reads should not swing wildly between replicates. If one replicate behaves differently, do not ignore it. Treat it as a signal that something changed in the experiment or sample handling.
For beginners, the most practical habit is comparison. Compare:
• ChIP vs Input
• Replicate A vs Replicate B
• Treatment vs Control under the same processing rules
This is where “strict quality inspection in every link” becomes real value. When QC is applied the same way each time, you can identify the true outlier quickly and avoid building conclusions on an unstable sample.
✓ Validate the dataset before peak calling saves time later
✓ Consistent filtering prevents artificial differences
✓ Outliers should be documented, not hidden
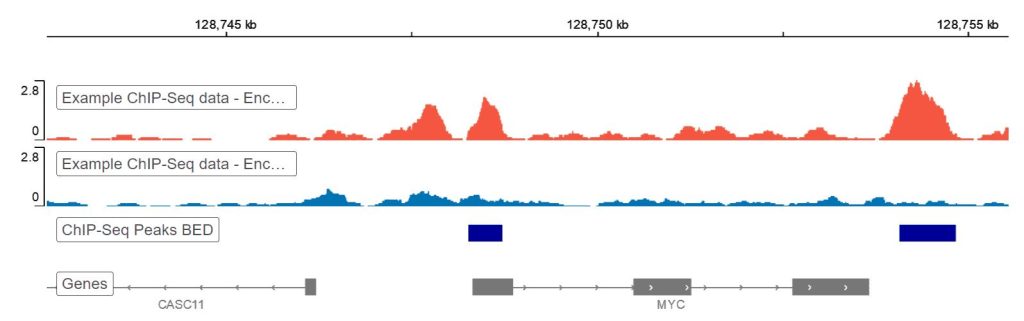
ChIP-Seq Analysis Tutorial - Basepair
4) Validate Peaks With Controls and Target-Appropriate Logic
Peak calling is not a single button. It is a decision system. A ChIP-seq target can be a narrow-binding transcription factor or a broader histone modification, and your validation method must match the biology.
Controls are the first proof layer. Input DNA helps model background signal and reduces false enrichment from open chromatin, repetitive regions, or sequencing bias. If peak sets change dramatically when controls are applied, you may be seeing background-driven peaks rather than true binding.
Next, validate peak shape and enrichment logic. Transcription factor peaks often appear sharp and localized, while some histone marks form broader regions. A mismatch between expected signal shape and observed tracks is a reason to re-check upstream experimental conditions, not just adjust software thresholds.
Longlight also supports analysis of specific genes or regions based on the customer’s research purpose. For many projects, this is the most actionable validation step. If your hypothesis focuses on a defined pathway or a target genomic region, validation should include targeted review of those loci, not only a global peak count.
✓ Use Input to separate enrichment from background
✓ Match peak strategy to TF vs histone behavior
✓ Validate key genes or regions that matter to your research question
5) Validate Reproducibility Before Any Differential Claim
The most persuasive validation is reproducibility. If replicates do not agree, “differential peaks” can become a story about noise rather than biology.
A practical ChIP-Seq Data Analysis Pipeline should check replicate agreement at two levels: genome-wide signal similarity and peak overlap stability. You can also examine whether top-ranked peaks remain top-ranked across replicates. If only one replicate drives the result, your conclusion is fragile.
For beginners, it helps to set a simple standard: you should be able to explain why a peak is considered reliable. That explanation usually includes consistent presence across replicates, stronger enrichment than control, and a signal profile that fits the target type.
This is also where strict QC helps customers in a direct way. Strong quality control reduces repeat experiments, and it improves the odds that your final report supports a publishable conclusion rather than an internal-only observation.
✓ Replicate agreement is a requirement, not a bonus
✓ Differential analysis should be built on stable peaks
✓ “Explainable peaks” are safer than “more peaks”
6) Turn Validation Into a Report You Can Use and Share
Validation is only valuable if it becomes a clear report that others can trust. A complete output should include raw data delivery, standardized QC summaries, peak calling settings, and the logic used to confirm reproducibility. It should also translate results into biological interpretation: peak annotation, target-region review, and meaningful examples that connect to your hypothesis.
Longlight Technology supports modern genomics with integrated solutions—NGS-related instruments, reagent consumables, and laboratory workflows used in academic, clinical, and industrial settings. In practice, this means researchers can align experimental execution with downstream analysis standards, instead of treating them as separate worlds. We also provide widely used consumables and kits such as Qubit tubes, nucleic acid extraction kits, and library preparation kits, which support stable workflows across multiple project types.
CTA (Call-To-Action): If you want a ChIP-seq project that is validated step-by-step—from sample handling to a complete data report—contact Longlight Technology to request a technical consultation and a free quote. We deliver a purpose-built validation checklist for your target category and ensure controls and reporting are agreed pre-sequencing for a smooth downstream workflow.
• Request an integrated ChIP-seq plan aligned to your research goals
• Nominate priority genes/regions for targeted analysis
• Obtain full reports with raw data and validation summaries